
Models
San Francisco, California
ZUNA is a 380M-parameter BCI foundation model for EEG data, a significant milestone in the development of noninvasive thought-to-text. ZUNA reconstructs, denoises, and upsamples EEG data across arbitrary channel layouts and is built for researchers, clinicians, and BCI developers using real world data.
Chris Warner, Jonas Mago, Jonathan Huml, Beren Millidge
Introduction
Zyphra is excited to announce ZUNA, our first foundation model trained on brain data. We believe thought-to-text will be the next major modality beyond language, audio, and vision enabled by noninvasive brain–computer interfaces (BCIs).
ZUNA is an early effort to build general foundation models of neural signals that can be used to understand and decode brain states. ZUNA is a key component in our mission to build human-aligned superintelligence. Over time, we see these models forming the foundation of thought-to-text agentic systems.
ZUNA
ZUNA is a 380M-parameter diffusion autoencoder trained to denoise, reconstruct, and upsample scalp-EEG signals. Given a subset of EEG channels, ZUNA can:
Denoise existing EEG channels
Reconstruct missing EEG channels
Predict novel channel signals, given physical coordinates on the scalp
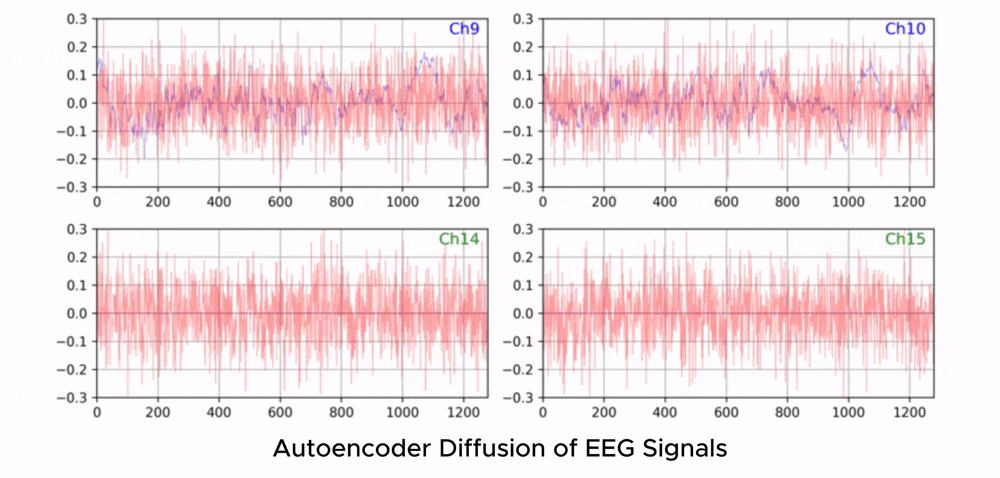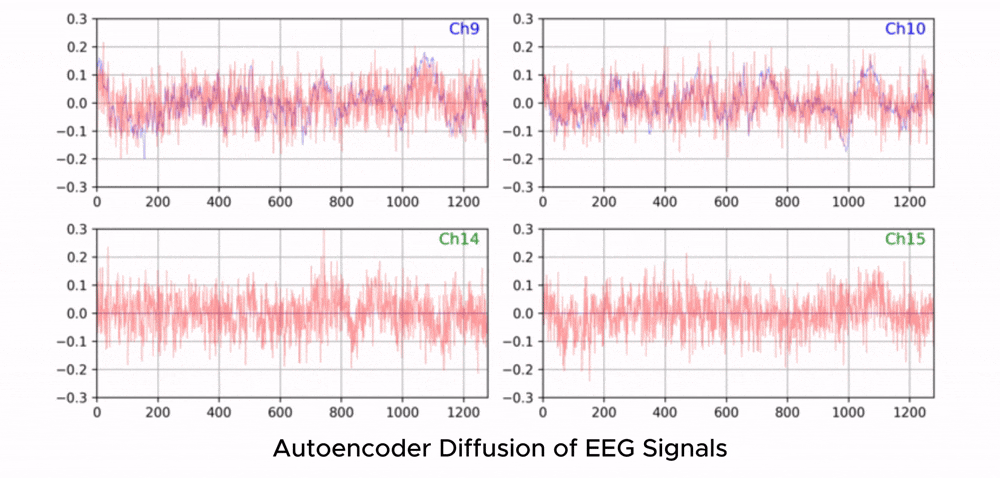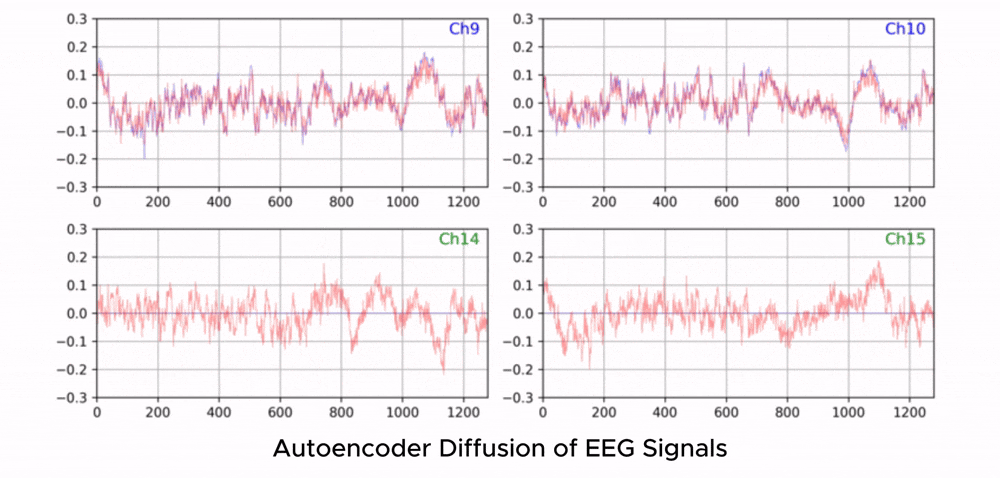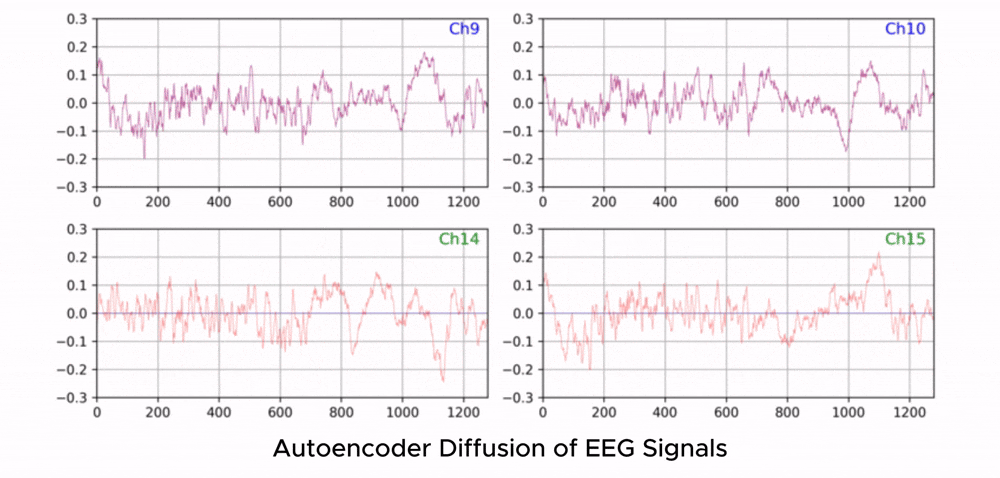
Why EEG Needs Foundation Models
EEG data is prevalent in clinics, research labs, and increasingly consumer devices. Yet unlike text, images, or audio, the EEG domain still lacks general and powerful foundation models.
Data fragmentation is a major reason for the lack of EEG foundation models. EEG datasets are typically small, collected under different protocols, and distributed across many institutions. This makes it difficult to aggregate data at the scale that has powered progress in other modalities.
And yet, there is clearly immense information and structure contained in EEG signals, which could power downstream tasks like understanding emotional and attentional states to decoding thoughts and dreams.
We aim to apply the classical deep learning methodology of creating general pretrained foundation models upon this data, with the goal of discovering generalizable representations underlying these signals and developing a foundation model that will translate thought-to-text.
Limitations with EEG Signal Processing
EEG recordings are frequently degraded by channel dropouts, motion-related artifacts, and limited channel counts typical of academic and consumer-grade hardware.
The standard approach for handling missing or noisy channels is spherical spline interpolation, which is the default method in the widely-used MNE package1. While simple and fast, this method has only a surface-level understanding of EEG structure and can result in poor or misleading reconstructions especially as channel degradation increases.
ZUNA replaces spherical spline interpolation with a learned, data-driven approach. By leveraging representations learned across a large and diverse EEG corpus, ZUNA can reconstruct signals in a way that captures the underlying patterns in brain activity rather than simple spatial smoothing.
This is not just exploratory research; it solves concrete, everyday problems faced by anyone working with EEG.
While other EEG foundation models have been published, we found few that had released model weights or usable code in a way that let us produce fair baselines for comparison.
Overcoming Limitations with ZUNA
ZUNA is a foundation model purpose-built to address the most persistent and costly limitations in EEG research and development. ZUNA enables the following capabilities:
Rescue and Reuse Existing Data
EEG datasets often contain sessions that are partially unusable due to corrupted channels or intermittent dropout. These sessions are frequently discarded, reducing sample size and statistical power. ZUNA enables researchers to recover usable signals from such recordings, effectively increasing dataset size without additional data collection.
Upgrade Low-Channel and Consumer Hardware
Many modern EEG devices trade spatial resolution, the number of electrodes on the device, for accessibility. ZUNA allows low-channel systems to be mapped into a higher-resolution signal space, narrowing the gap between consumer-grade and lab-grade recordings and enabling analyses that would otherwise be infeasible.
Reduce Dependence on Fixed Electrode Montages
Traditional EEG analysis pipelines assume fixed montages (e.g., 10–20 or 10–10). ZUNA operates directly on electrode coordinates, allowing it to generalize across arbitrary channel counts and layouts. This makes cross-dataset and cross-device analyses substantially easier.
Results
We compared the performance of ZUNA to spherical spline interpolation which is ubiquitously used by EEG researchers and practitioners and is included as the default in the widely-used MNE package. Spherical spline interpolation is a surprisingly robust baseline when relatively small numbers of channels are dropped out, but degrades in performance with increasingly degraded data. We evaluated ZUNA’s performance on a validation set, a portion of our training data the model had not seen, and on several unseen test datasets of different distributions.
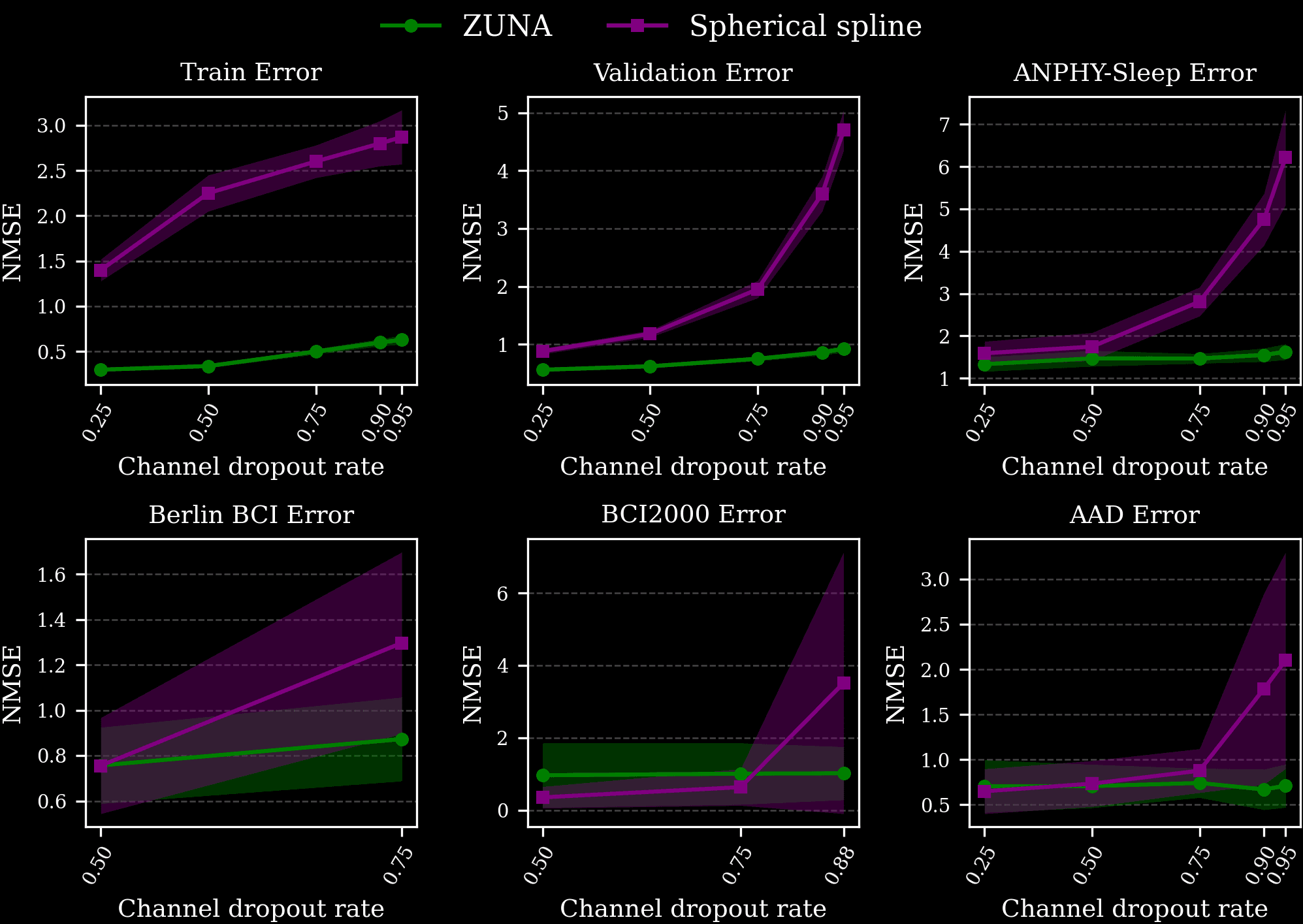
We find that across datasets, ZUNA outperforms spherical spline interpolation by a significant margin, with the advantage increasing at high levels of dropout. When dropping more than 75% of channels, ZUNA outperforms spherical spline interpolation across all datasets.
Architecture
ZUNA leverages a diffusion autoencoder architecture based on a transformer backbone. An encoder maps EEG signals to a shared latent space and a decoder reconstructs EEG signals from latents. We trained with a masked reconstruction loss and a heavy dropout scheme, enabling ZUNA to denoise existing channels and predict new ones during inference.
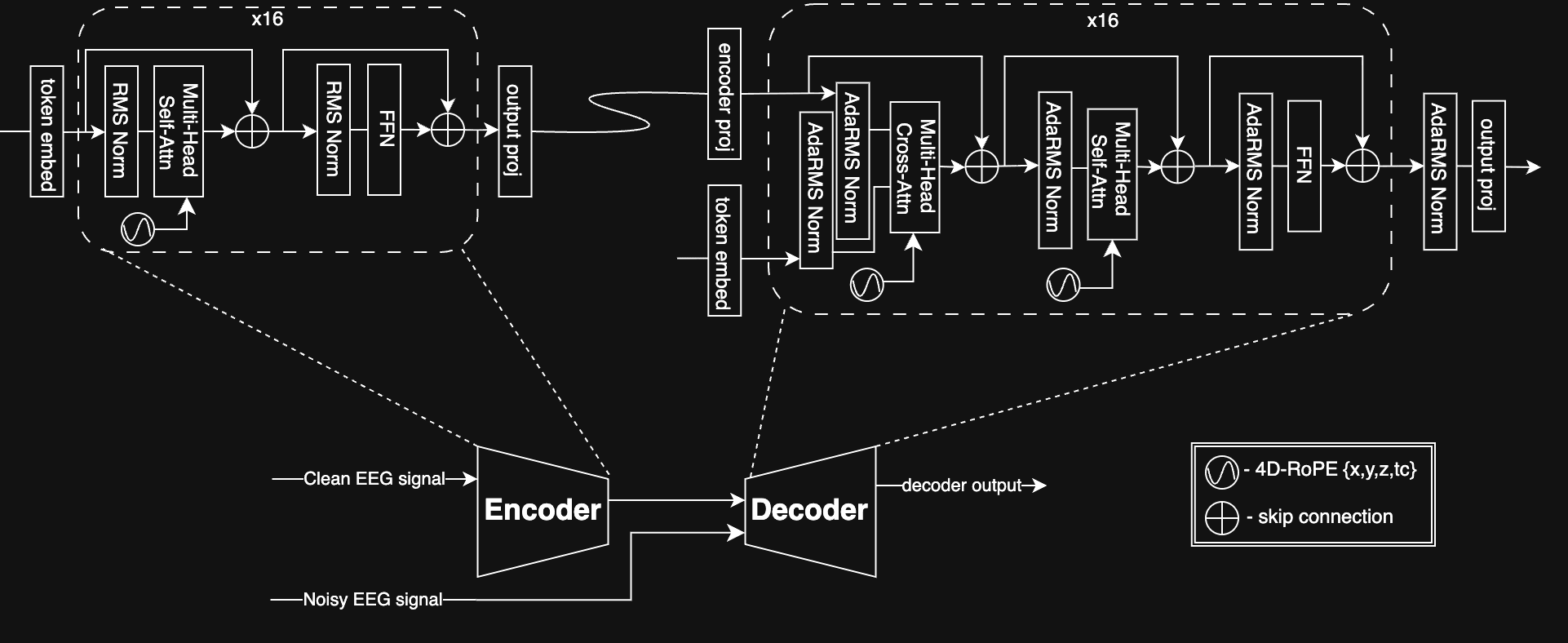
To handle EEG data, which can contain an arbitrary number of channels in arbitrary positions on the scalp, we introduce two architectural innovations:
To adapt the transformer architecture to a heterogeneous number of channels where each channel is a continuous, real-valued signal, we first compressed each channel’s signal into 0.125s ‘chunks’ which the model learned to map to continuous ‘tokens’. We then rasterized these tokens into a single 1-D sequence that can be operated upon by the standard transformer architecture.
To represent the physical location in space of the recording electrodes, which can differ across datasets or even samples, we utilized 4-D RoPE as position embeddings. For each channel token we encoded the electrode x, y, z positions and the coarse time dimension as separate into separate components of the attention head dimension. We found that this encoding method enabled us to efficiently generalize to novel x, y, z positions in a smooth and stable way while still representing the sequence ordering across time.
Data
To train ZUNA, we curated approximately 2 million channel-hours of EEG data from a wide range of publicly available sources. All data used a standardized preprocessing pipeline to make it suitable for large-scale foundation model training.
We plan to open-source our data and data infrastructure. Large, high-quality public datasets are essential for progress in any deep learning domain, EEG included.
Release
We are releasing ZUNA with a permissive licensing (Apache 2.0) and practical tooling so it can be easily integrated into real-world workflows.
Model weights: ZUNA Hugging Face
Inference & preprocessing code: ZUNA GitHub
Pip install Python package: pip install zuna
At 380 million parameters, ZUNA is lightweight enough to run quickly on a consumer GPU and can be used on CPU for many workloads.
We hope researchers, clinicians, and builders put ZUNA to work, provide feedback, and help shape the next generation of brain foundation models.
Organizations or researchers interested in collaborating with Zyphra to improve future versions for specific needs or use cases should contact us at bci@zyphra.com.
Disclaimer: This website and related services (“Services”) are provided for research use only and is not intended for use in the diagnosis, cure, mitigation, treatment, or prevention of any disease or health condition. The Services have not been validated for any medical or clinical use. The information provided through the Services is for general informational purposes only and is not a substitute for any professional medical or healthcare advice. We do not warrant that any information provided through the Services is accurate, complete, or useful to you. Any reliance you place on such information is strictly at your own risk.